01/3D Structure
? About the 3D Viewer
Mol* (pronounced "molstar") is an open-source molecular visualization tool used by the Protein Data Bank and AlphaFold Database. Learn more at molstar.org.
Controls:
- Rotate: Click and drag
- Zoom: Scroll wheel or pinch
- Pan: Right-click and drag (or two-finger drag)
- Reset: Double-click to reset view
What am I looking at?
This is a predicted 3D structure of the protein. The ribbon diagram shows the protein backbone—helices appear as coils, sheets as arrows, and loops as simple lines. The shape determines how the protein functions: where it binds to other molecules, how it catalyzes reactions, and how mutations might disrupt its activity.
Color legend:
The structure is colored by pLDDT confidence score, which indicates how confident AlphaFold is in each region's predicted position:
- Blue (>90): Very high confidence
- Cyan (70-90): Confident
- Yellow (50-70): Low confidence
- Orange (<50): Very low confidence, likely disordered
02/AI Analysis
TLDR
The TAU protein helps stabilize the internal scaffolding of brain cells, and the P301L mutation causes a devastating inherited form of dementia by making this protein clump together abnormally. Scientists used AI-based structure prediction to study how P301L alters TAU's shape, but the resulting model had very low confidence (average score 55 out of 100), indicating the protein exists in flexible, constantly changing forms rather than a single stable structure. This flexibility is actually scientifically important—it helps explain why P301L accelerates the formation of toxic protein aggregates that kill neurons in frontotemporal dementia and related conditions.
Detailed Analysis
Works Cited
Similar Research
03/Research Data
ClinVar Classification
Not found in ClinVar
Population Frequency
No population data available
Disease Associations
1182 totalShowing 5 of 1182 associations
AI Research Brief
Research brief will be generated when agent findings are available.
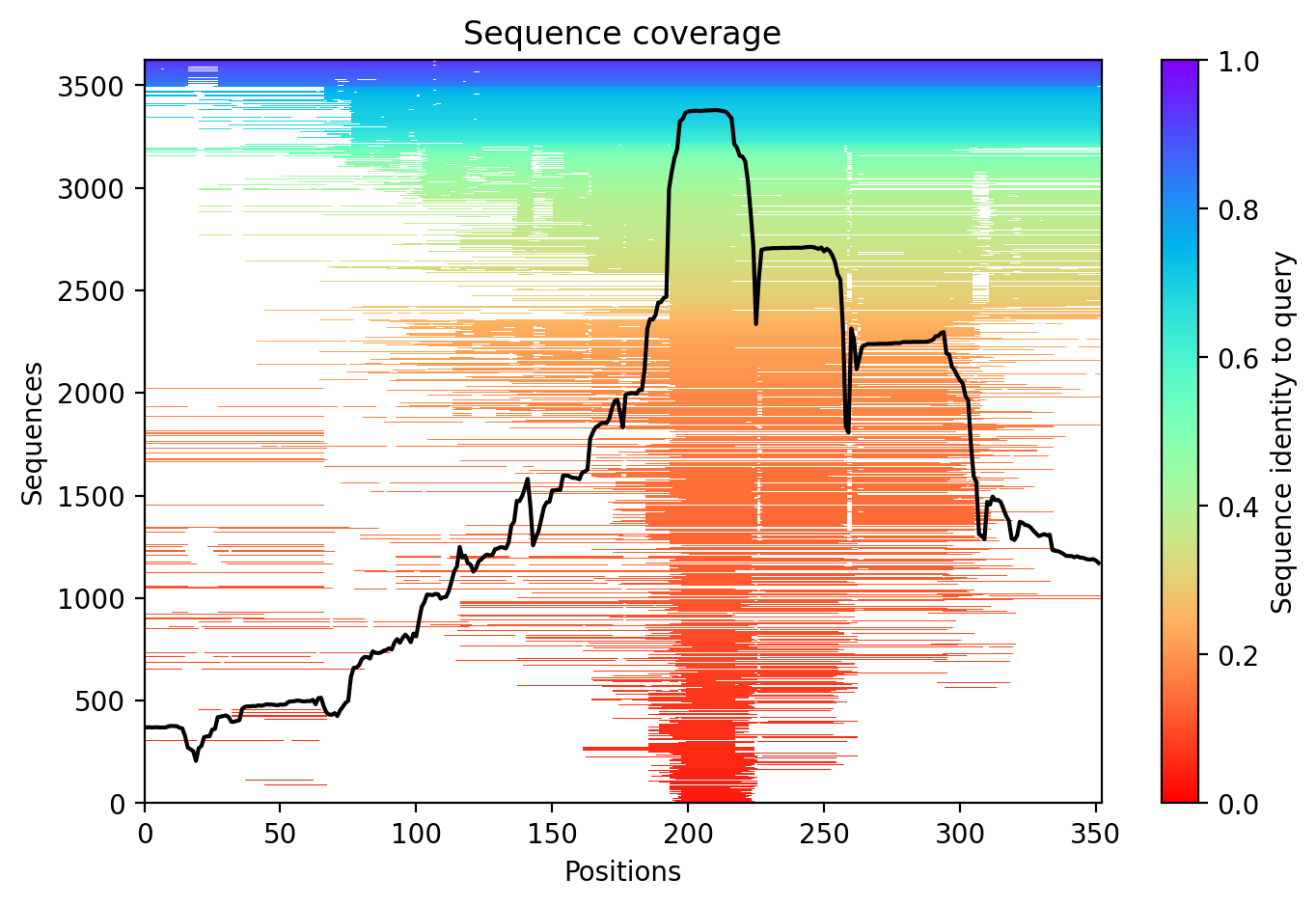
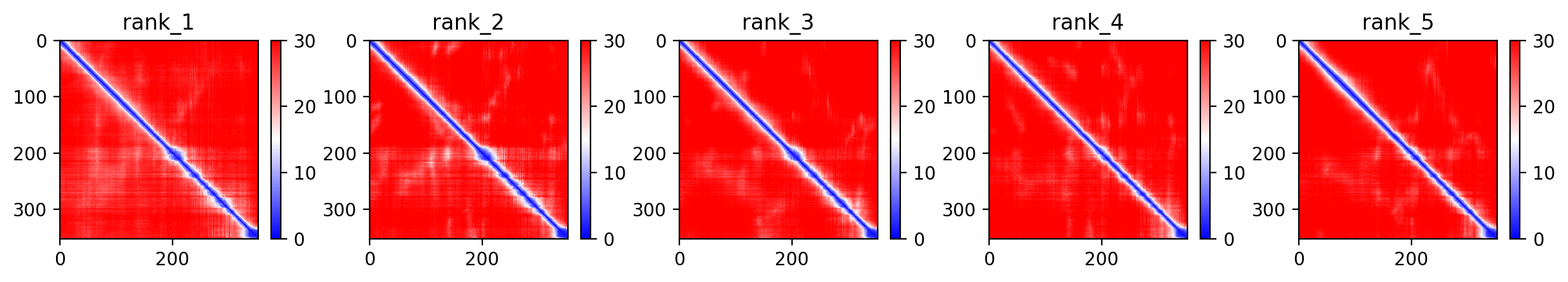
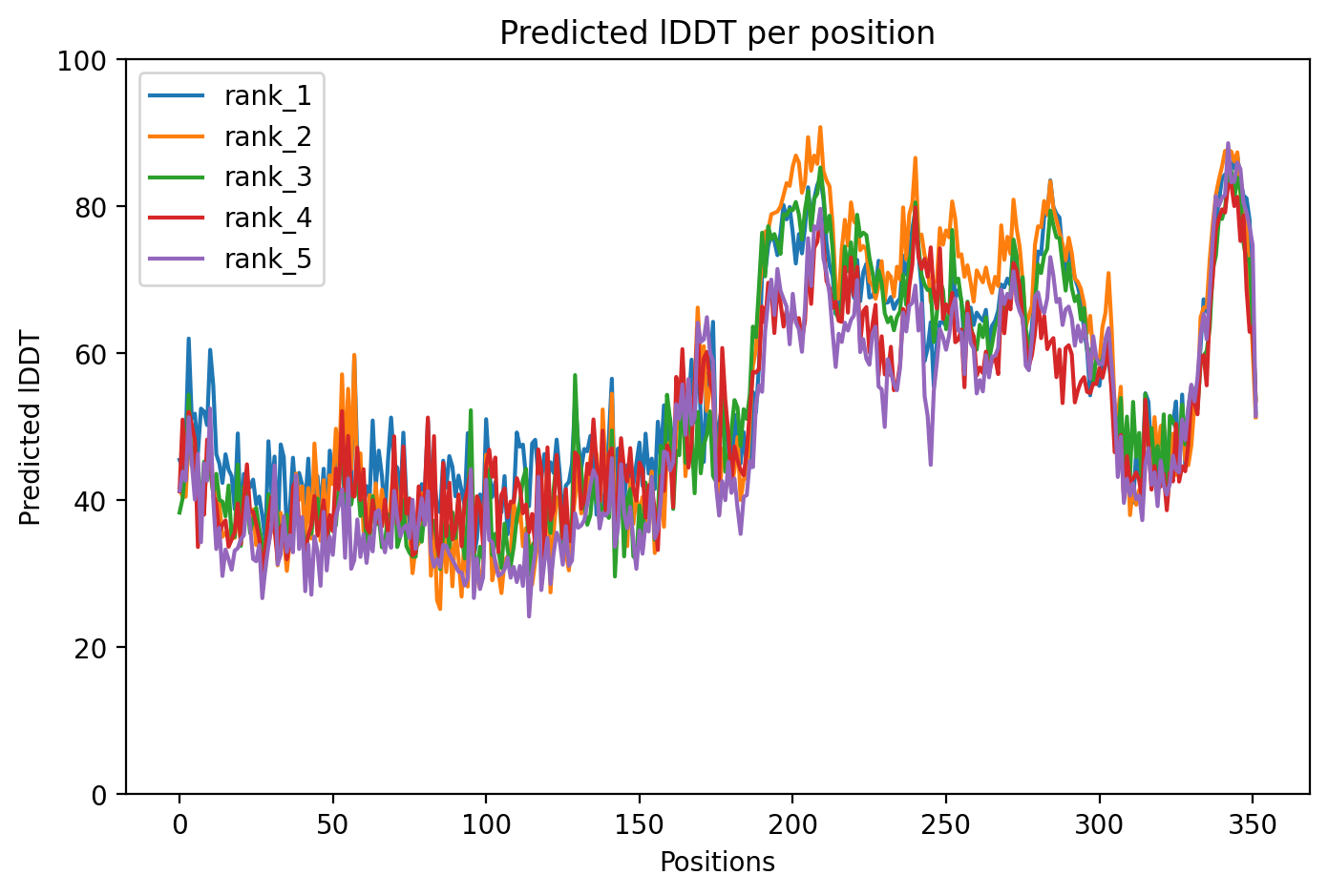
04/AlphaFold Metrics
05/Domain Annotations
Structural Domains & Regions
Binding Partners
Gene Ontology
06/Structural Caption
TAU P301L variant shows intrinsically disordered architecture (average pLDDT 55.0) with structured microtubule-binding repeats, where mutation promotes aggregation-prone conformation.
Average pLDDT of 55.0 with only 19% high-confidence residues (67/352) indicates a largely intrinsically disordered protein. The microtubule-binding domain (residues 561-685) shows the highest structural confidence, while N-terminal and C-terminal regions remain destabilized.
The four tandem Tau/MAP repeats (residues 561-685) constitute the microtubule-binding domain and represent the most structured region. Extensive disordered regions (residues 1-573, 715-734) and low complexity segments correlate with low pLDDT throughout most of the protein, consistent with Tau's known intrinsically disordered character.
The P301L mutation in the microtubule-binding domain disrupts proline-mediated structural constraints, potentially accelerating pathological aggregation and reducing microtubule-binding affinity characteristic of frontotemporal dementia-associated Tau variants.
07/Peptide Therapeutics
Aggregation analysis pending. Run peptide agent to compute aggregation propensity.
08/Known Inhibitors
No known inhibitors found. Run peptide agent to search literature.
09/Candidate Peptides
No candidate peptides generated yet. Run peptide agent to design inhibitory peptides.
10/Agent Findings
No agent findings yet. Research agents analyze folds on scheduled intervals.
11/Agent Annotations
No agent annotations yet. Agents can submit annotations via the API.